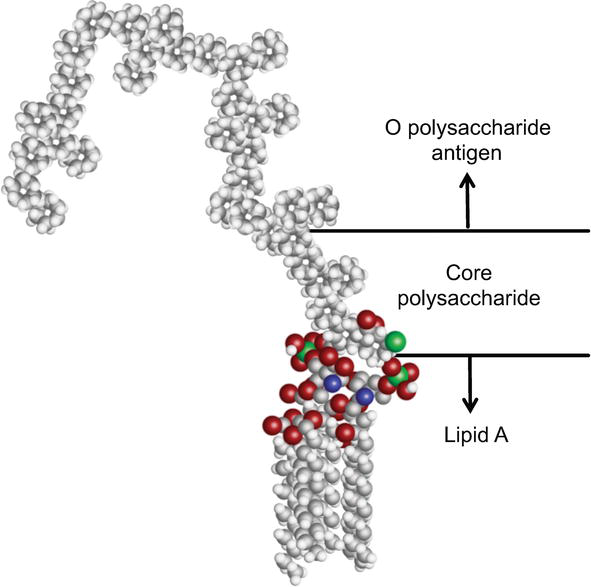
Lipopolysaccharide Structure and Biosynthesis in Helicobacter pylori - Li - 2016 - Helicobacter - Wiley Online Library

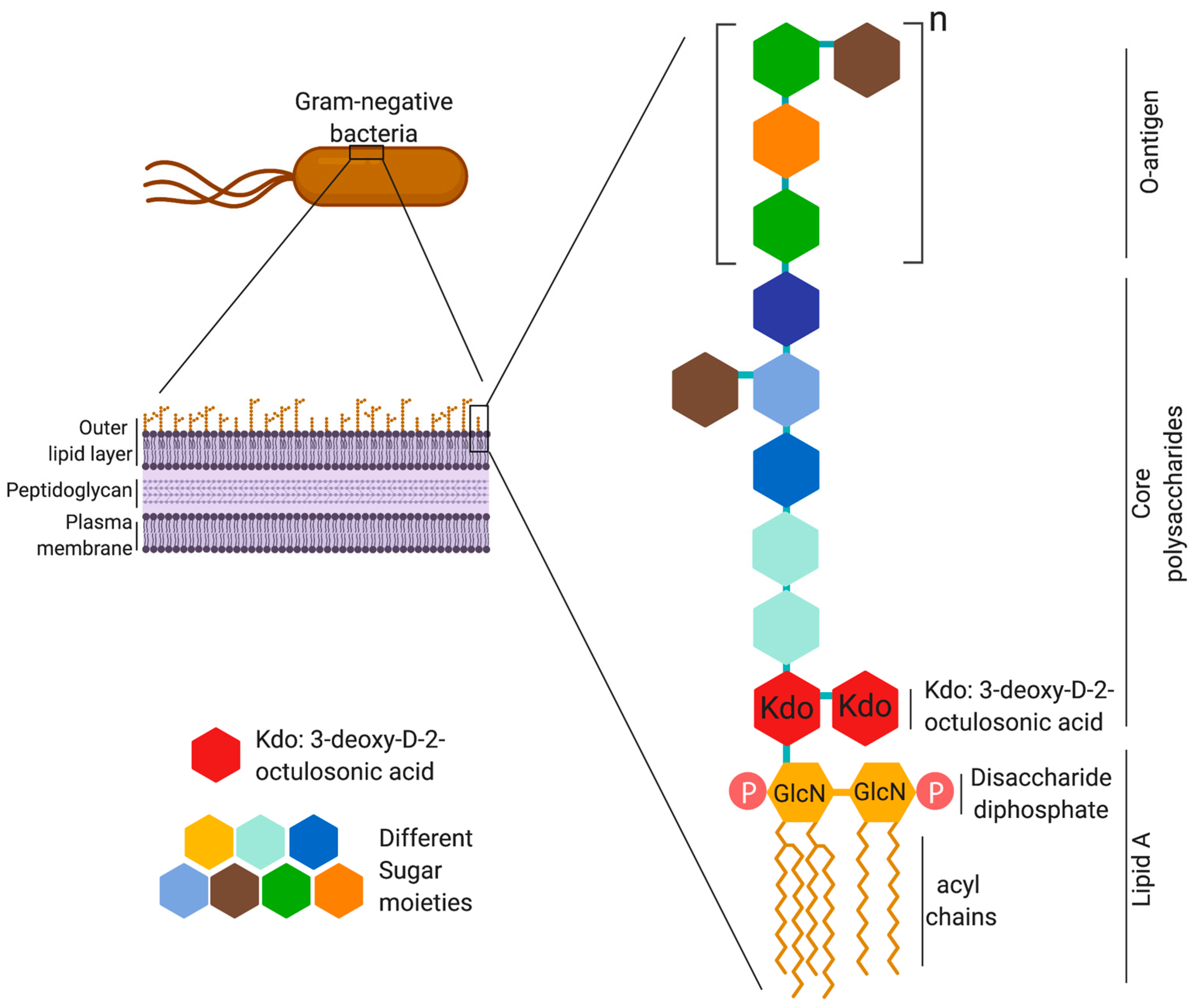
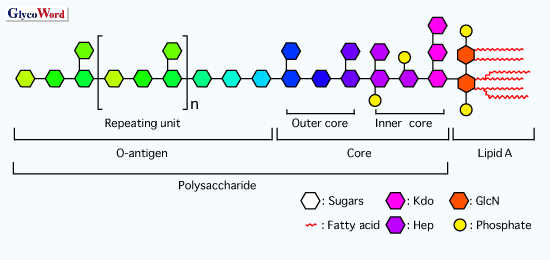
Schematic of the basic structure of lipopolysaccharide. LPS consists of... | Download Scientific Diagram

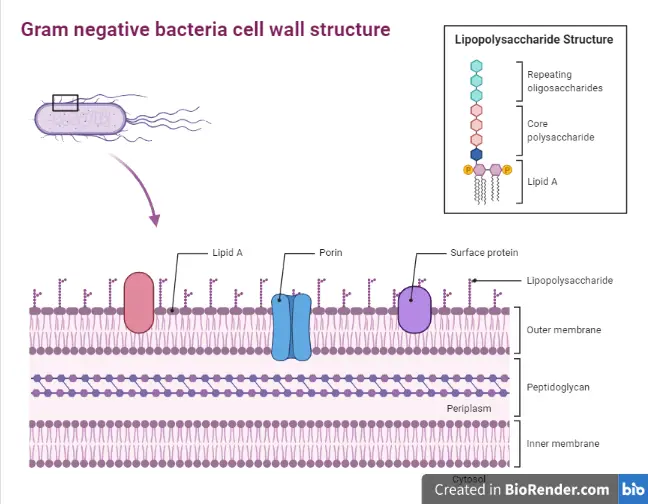
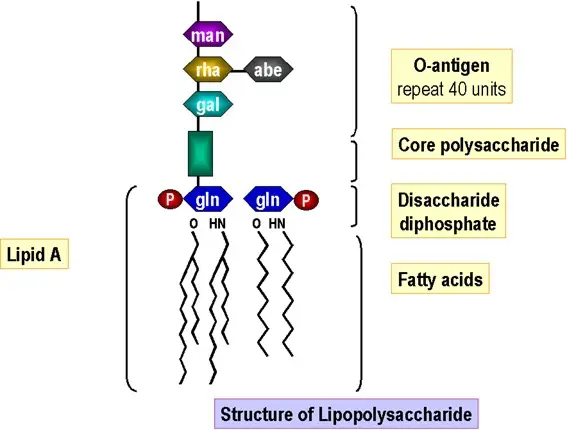
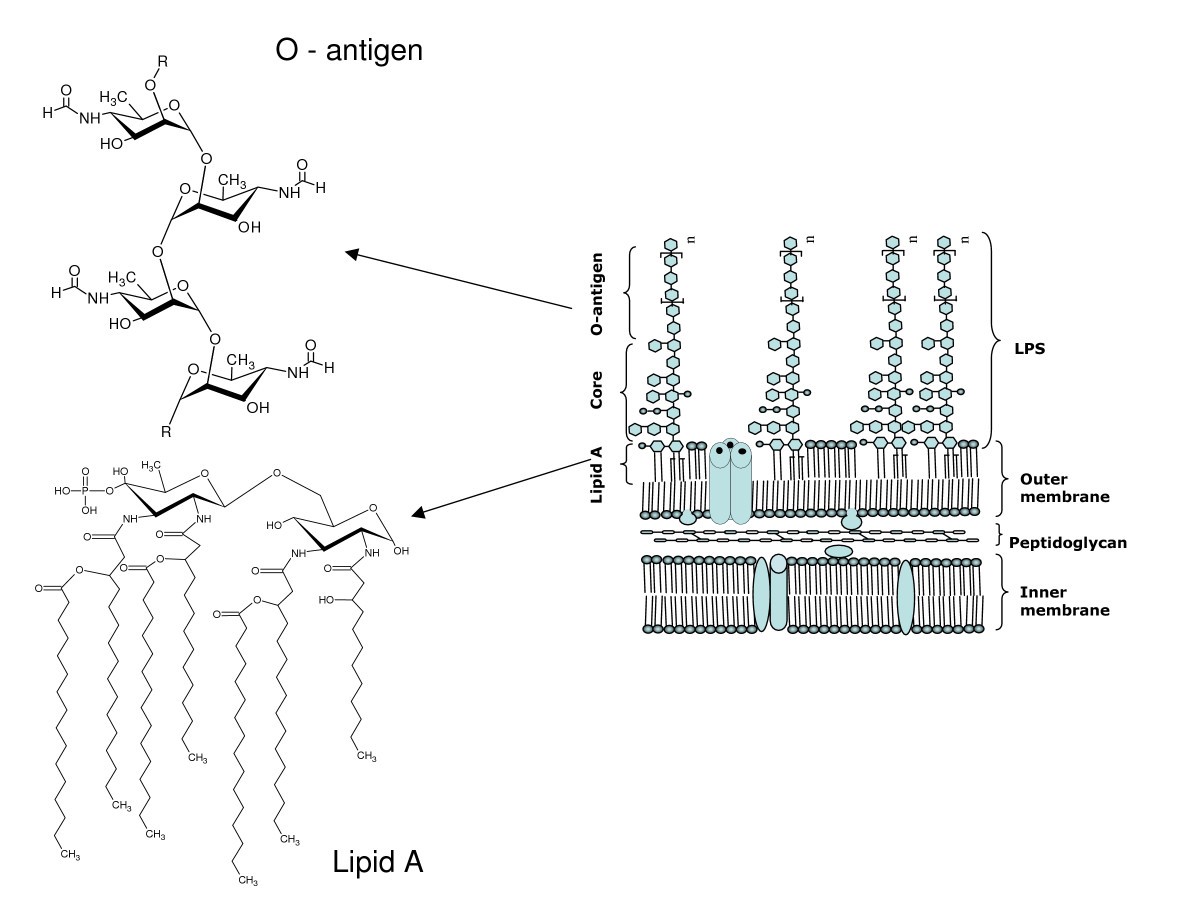
Schematic representation of the structure of lipopolysaccharide (LPS). | Download Scientific Diagram

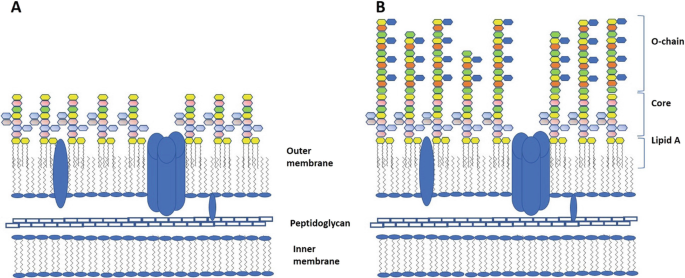
Structural Characterization of a Model Gram-Negative Bacterial Surface Using Lipopolysaccharides from Rough Strains of Escherichia coli | Biomacromolecules
Brucella spp noncanonical LPS: structure, biosynthesis, and interaction with host immune system | Microbial Cell Factories | Full Text

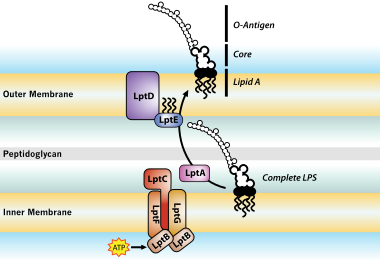
LptE binds to and alters the physical state of LPS to catalyze its assembly at the cell surface | PNAS

Gram-negative trimeric porins have specific LPS binding sites that are essential for porin biogenesis | PNAS

Aggregation Behavior of an Ultra-Pure Lipopolysaccharide that Stimulates TLR-4 Receptors: Biophysical Journal
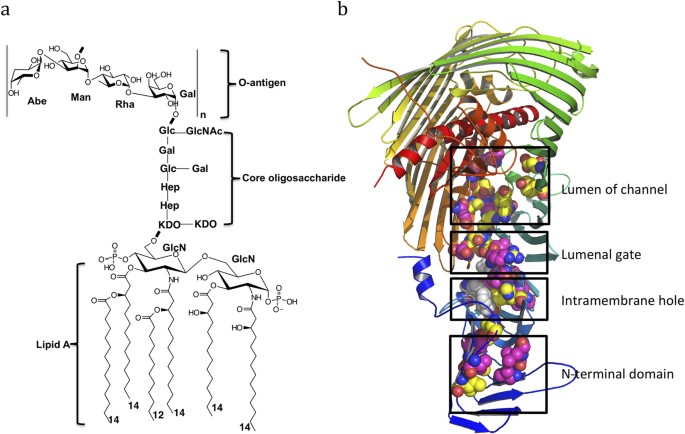
Trapped lipopolysaccharide and LptD intermediates reveal lipopolysaccharide translocation steps across the Escherichia coli outer membrane | Scientific Reports
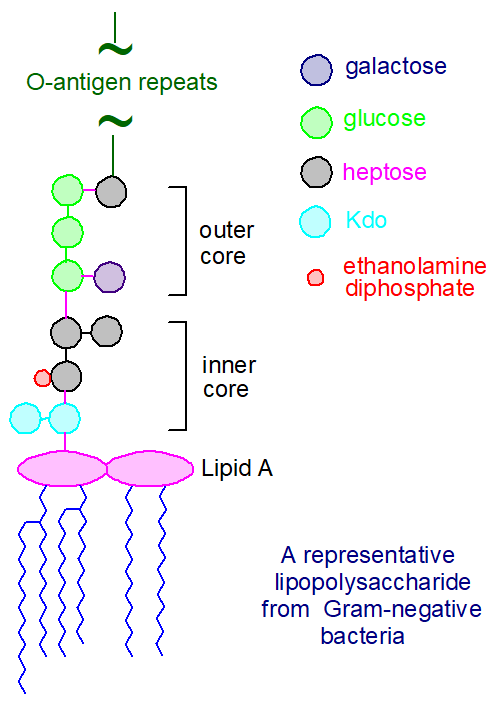
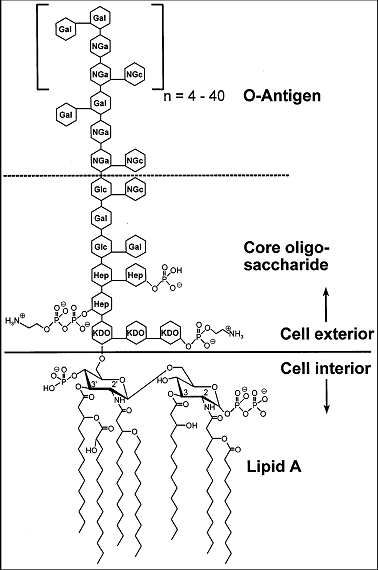
Lipid A and Bacterial Lipopolysaccharides, Gram-negative, endotoxins, lipo-chitooligosaccharides (Nod factors) - structure, occurrence, biochemistry and function

Pairing Bacteroides vulgatus LPS Structure with Its Immunomodulatory Effects on Human Cellular Models | ACS Central Science